Extending and comparing methods for synthetic social contact matrices
SPECTRUM & SPARK Annual Meeting
Nicholas Tierney
Telethon Kids Institute
15th Feb, 2023
A contact matrix
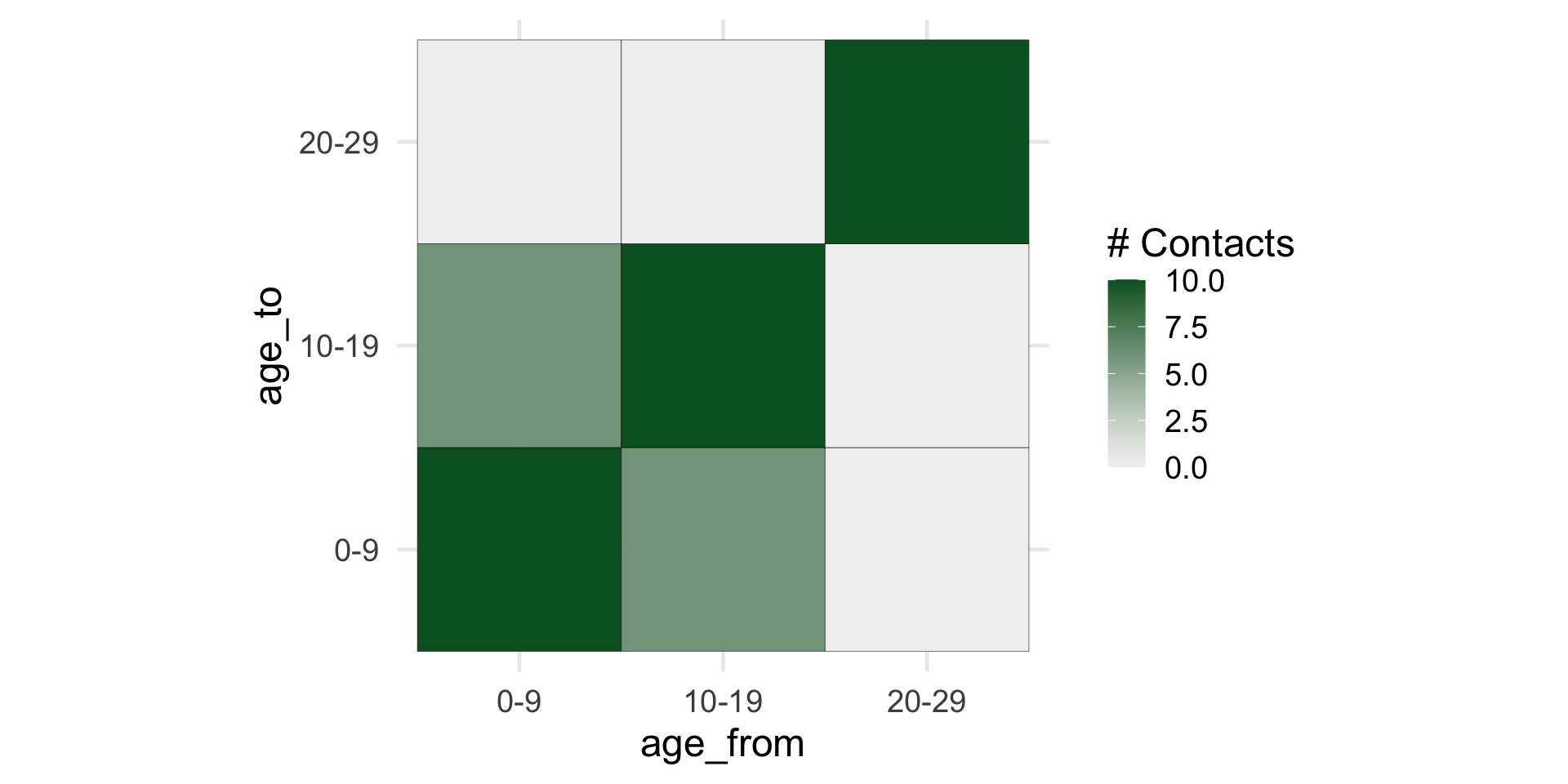
Mossong et al (POLYMOD)

Prem et al - Synthetic Contact Matrices

Populations are different
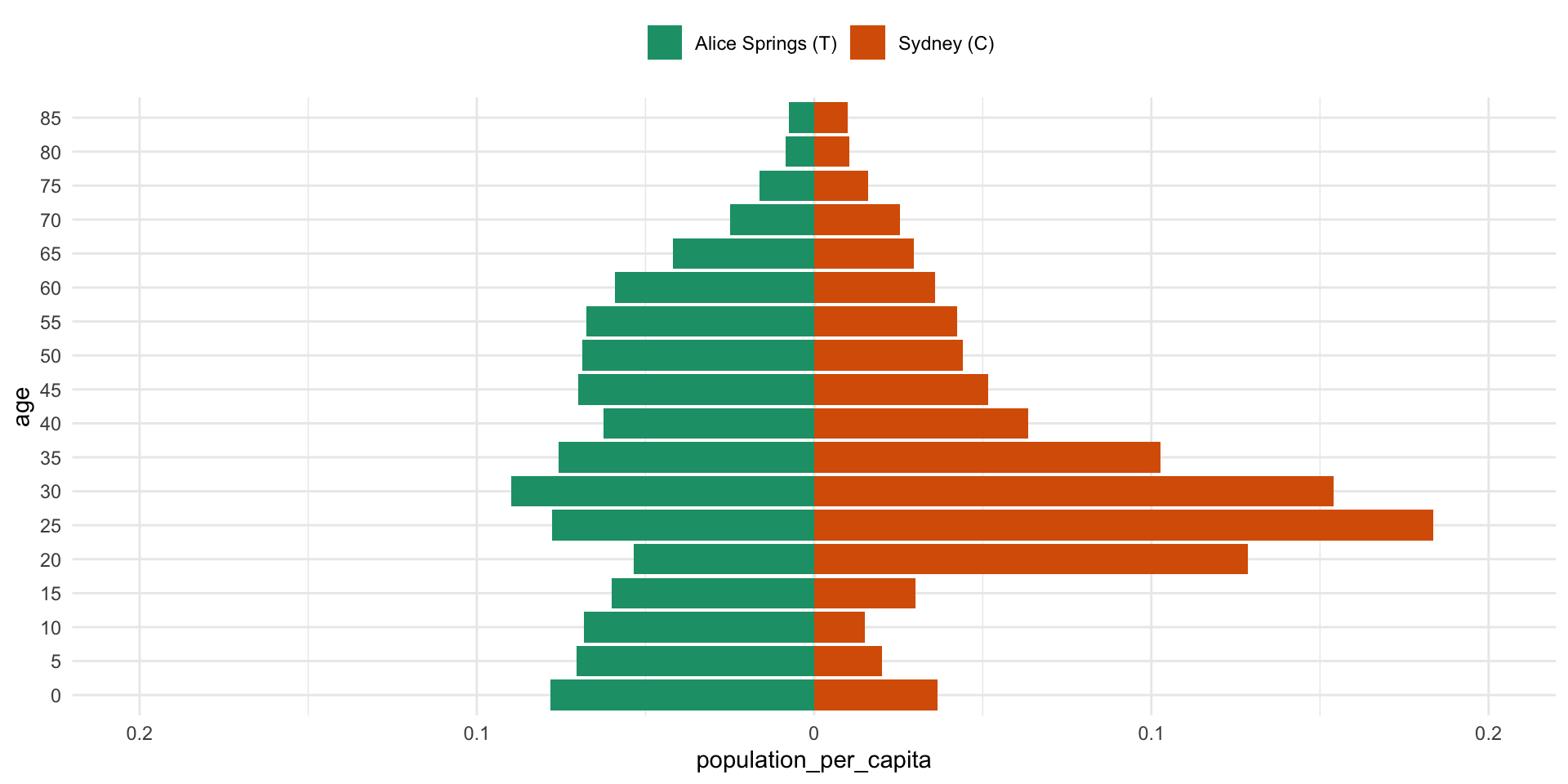
Key points
- Contact matrices can model infectious disease spread
- We need contact matrices for specific parts of Australia
- Current best estimates are for all of Australia
- We need our own approach
We need an R package: conmat
- Input: age population data
- Output: synthetic contact matrix
- Provide tools to:
- Create next generation matrices
- Apply vaccination
- Use these in SIR modelling
Age population data
# A tibble: 18 × 4 (conmat_population)
- age: lower.age.limit
- population: population
lga lower.age.limit year population
<chr> <dbl> <dbl> <dbl>
1 Fairfield (C) 0 2020 12261
2 Fairfield (C) 5 2020 13093
3 Fairfield (C) 10 2020 13602
4 Fairfield (C) 15 2020 14323
5 Fairfield (C) 20 2020 15932
6 Fairfield (C) 25 2020 16190
7 Fairfield (C) 30 2020 14134
8 Fairfield (C) 35 2020 13034
9 Fairfield (C) 40 2020 12217
10 Fairfield (C) 45 2020 13449
11 Fairfield (C) 50 2020 13419
12 Fairfield (C) 55 2020 13652
13 Fairfield (C) 60 2020 12907
14 Fairfield (C) 65 2020 10541
15 Fairfield (C) 70 2020 8227
16 Fairfield (C) 75 2020 5598
17 Fairfield (C) 80 2020 4006
18 Fairfield (C) 85 2020 4240Predict (extrapolate) a new contact matrix
── Setting Prediction Matrices ────────────
A list of matrices containing the model
predicted contact rate between ages in
each setting.
• home: a 16x16 <matrix>
• work: a 16x16 <matrix>
• school: a 16x16 <matrix>
• other: a 16x16 <matrix>
• all: a 16x16 <matrix>
ℹ Access each <matrix> with `x$name`
ℹ e.g., `x$home`Synthetic Contact matrix
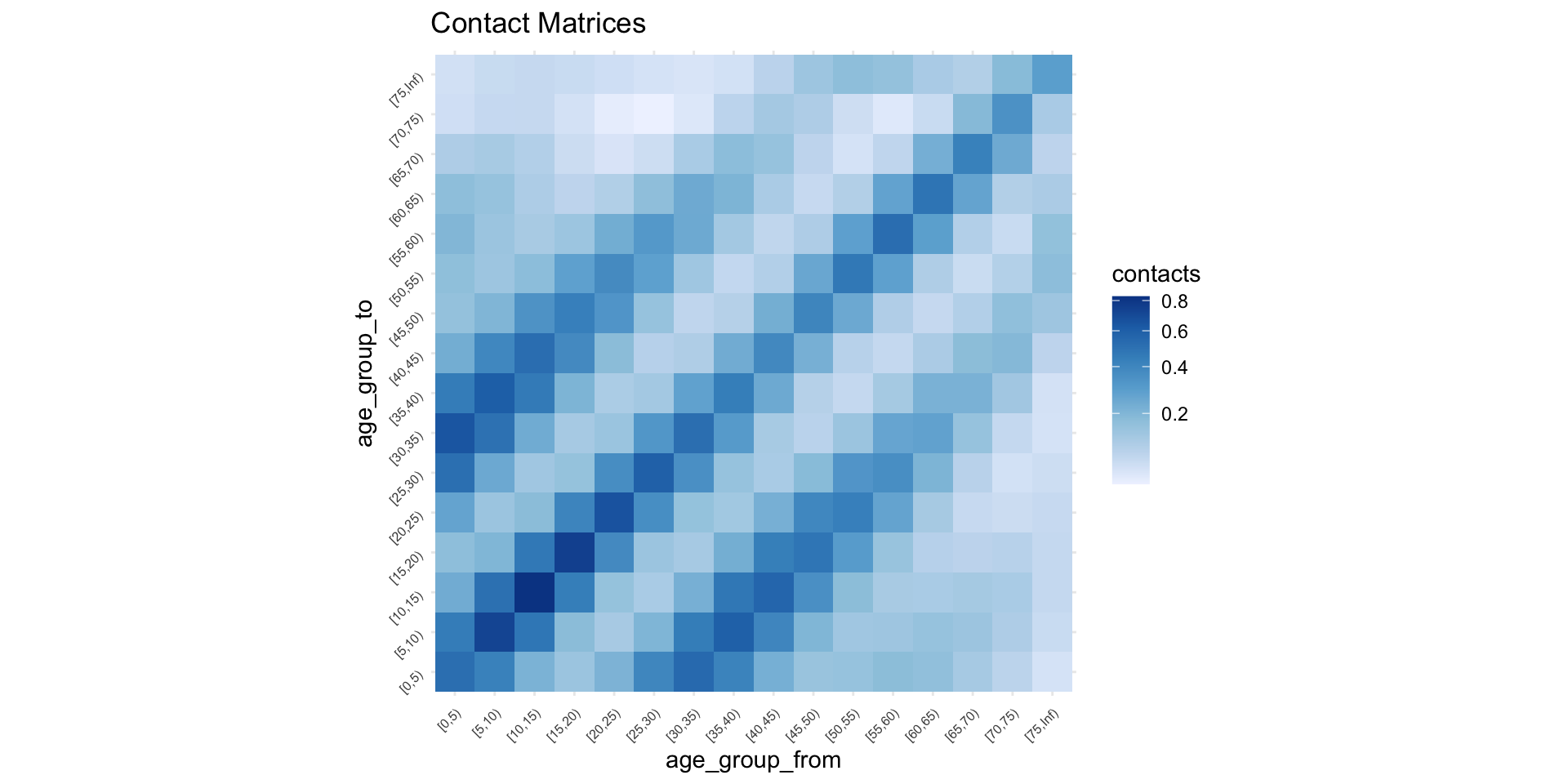
Next Generation Matrix
── NGM Setting Matrices ──────────────────
A list of matrices, each <matrix>
containing the number of newly infected
individuals for a specified age group.
• home: a 16x16 <matrix>
• school: a 16x16 <matrix>
• work: a 16x16 <matrix>
• other: a 16x16 <matrix>
• all: a 16x16 <matrix>
ℹ Access each <matrix> with `x$name`
ℹ e.g., `x$home`Next Generation Matrix
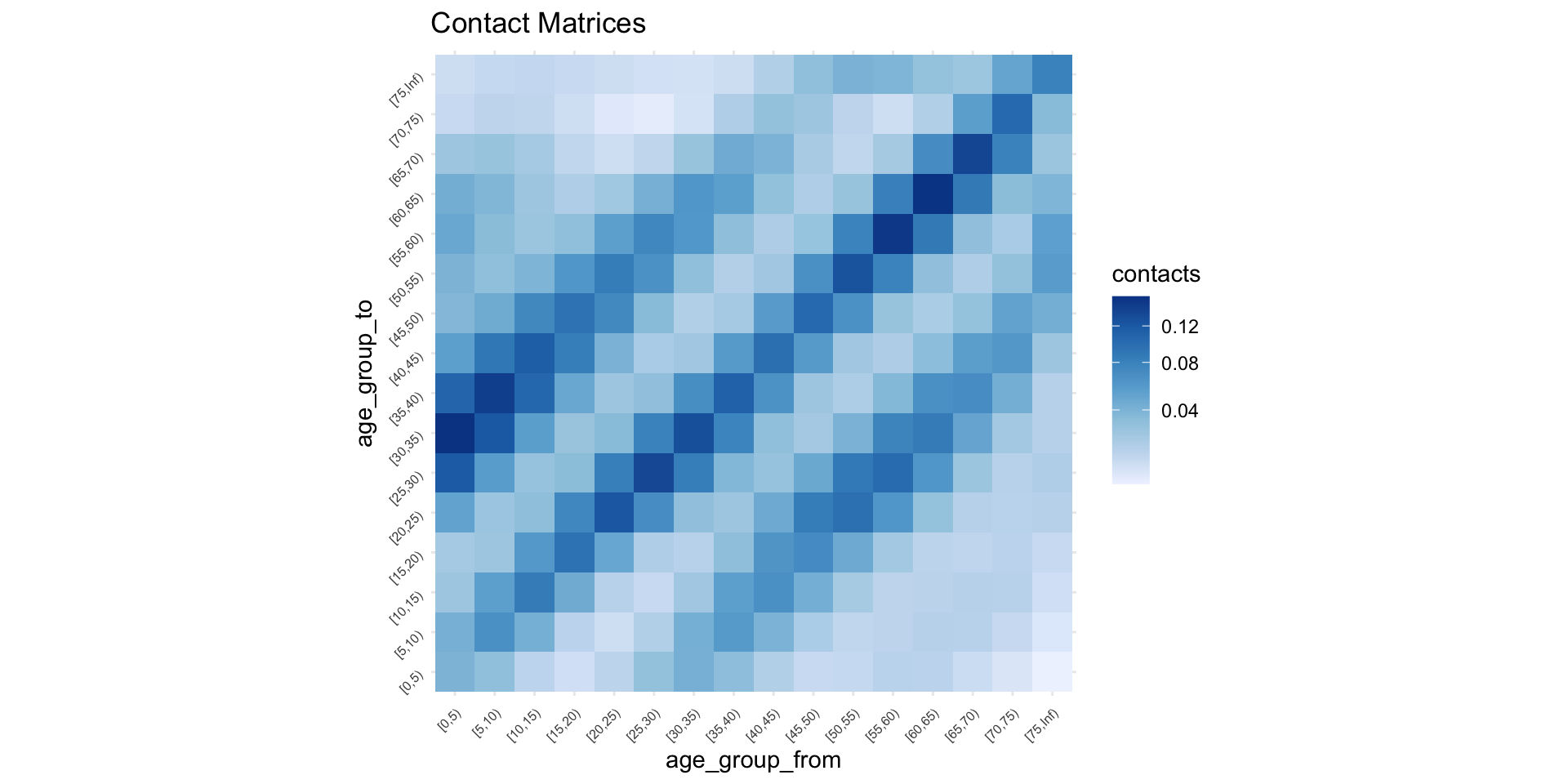
But is it different to Prem?
- Compare our contact matrix to Prem’s contact matrix
- In a Suscepible, Infected, Recovered (SIR) model
- For say, Germany?
…Is it different to Prem?
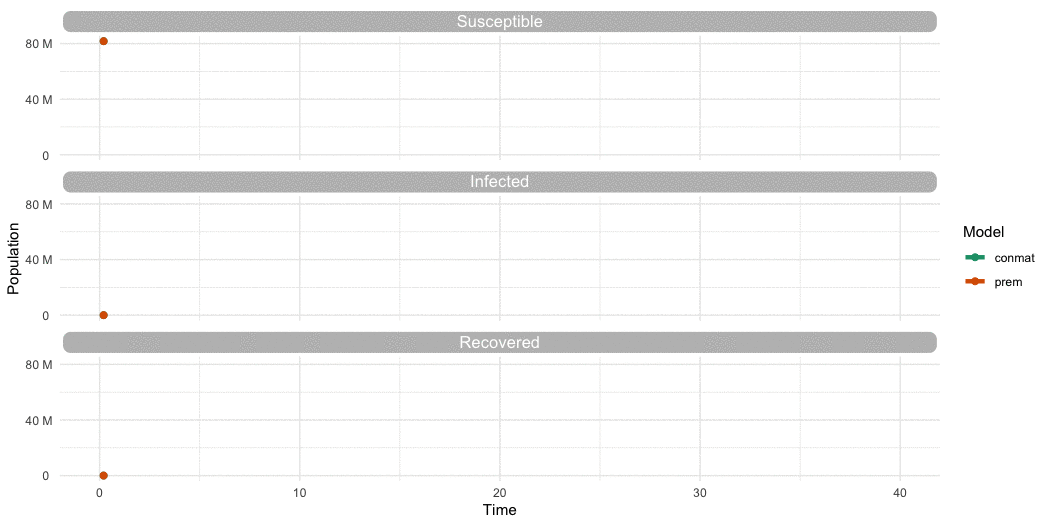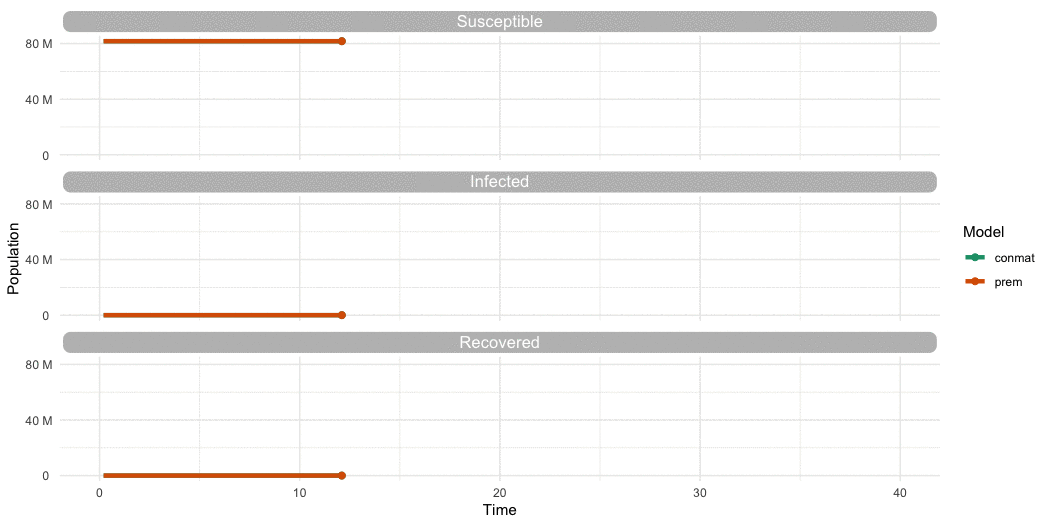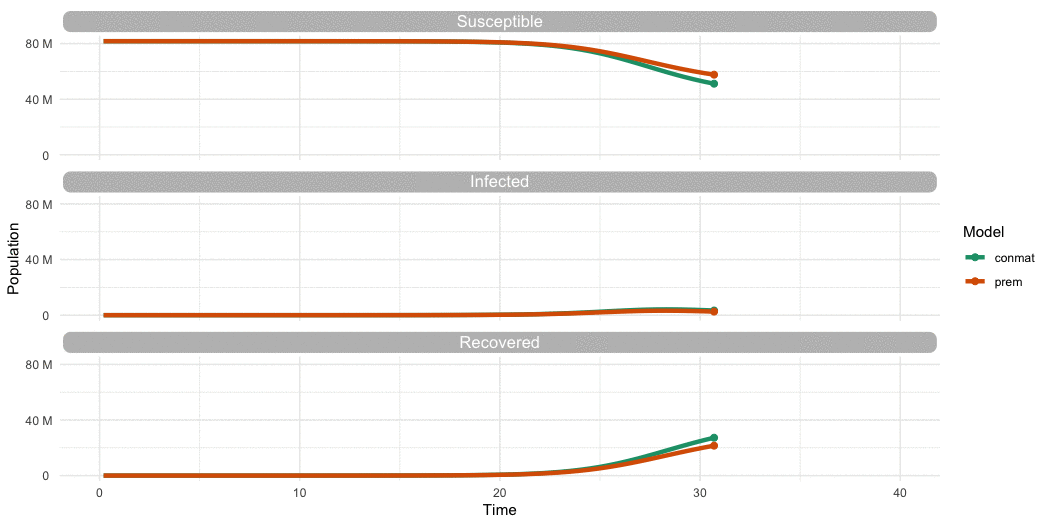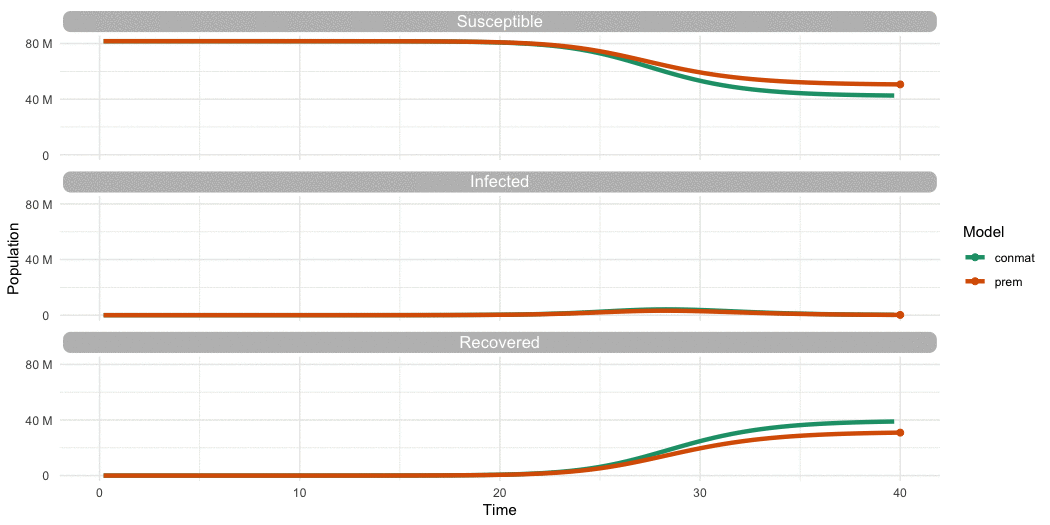
…Yes
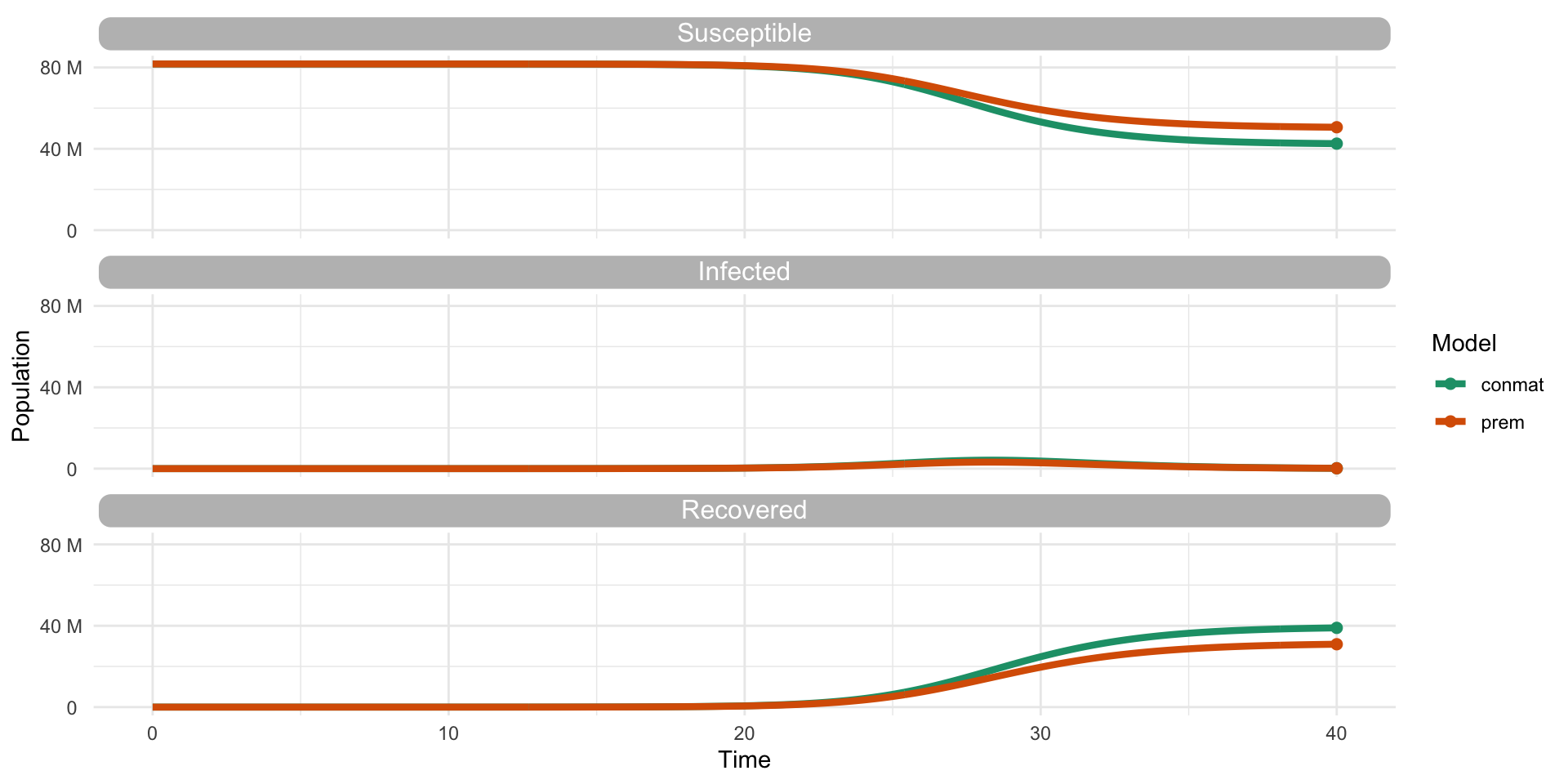
…Yes (but we need to look further)
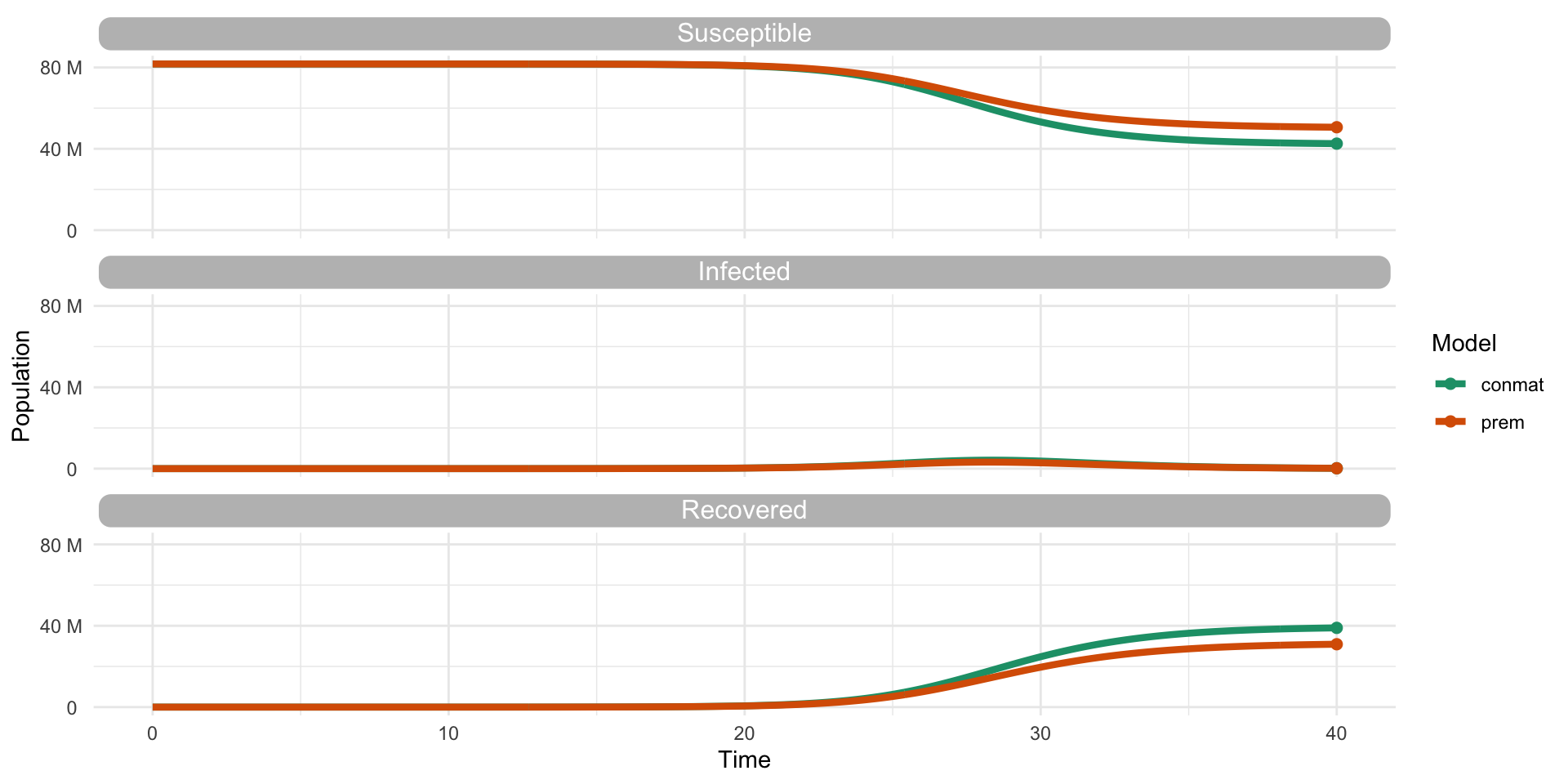
Thanks
- Nick Golding
- Aarathy Babu
- Michael Lydeamore
- Spark Spectrum Seed Funding
Learning more
github.com/njtierney/talk-spark-2023
njtierney@aus.social
nj_tierney
njtierney
nicholas.tierney@gmail.com
End.
Extras
Next Generation Matrices
- Once infected, a person can transmit an infectious disease to another, creating generations of infected individuals.
- We can define a matrix describing the number of newly infected individuals in age groups, for consecutive generations.
- This matrix is called a next generation matrix (NGM).
Apply Vaccination
Applies the effect of vaccination on the next generation of infections, to understand and describe the reduction of acquisition and transmission in each age group.
Takes the following arguments:
ngm- a Next Generation Matrixdata- A data frame with location specifics- which columns are related to “coverage”, “acquisition”, and “transmission” in the
datacol
Example vaccination data
# A tibble: 17 × 4
age_band coverage acquisition transmission
<chr> <dbl> <dbl> <dbl>
1 0-4 0 0 0
2 5-11 0.782 0.583 0.254
3 12-15 0.997 0.631 0.295
4 16-19 0.965 0.786 0.469
5 20-24 0.861 0.774 0.453
6 25-29 0.997 0.778 0.458
7 30-34 0.998 0.803 0.493
8 35-39 0.998 0.829 0.533
9 40-44 0.999 0.841 0.551
10 45-49 0.993 0.847 0.562
11 50-54 0.999 0.857 0.579
12 55-59 0.996 0.864 0.591
13 60-64 0.998 0.858 0.581
14 65-69 0.999 0.864 0.591
15 70-74 0.999 0.867 0.597
16 75-79 0.999 0.866 0.595
17 80+ 0.999 0.844 0.556Apply vaccination
── Vaccination Setting Matrices ──────────
A list of matrices, each <matrix>
containing the adjusted number of newly
infected individuals for age groups.
These numbers have been adjusted based on
proposed vaccination rates in age groups
• home: a 17x17 <matrix>
• school: a 17x17 <matrix>
• work: a 17x17 <matrix>
• other: a 17x17 <matrix>
• all: a 17x17 <matrix>
ℹ Access each <matrix> with `x$name`
ℹ e.g., `x$home`How does the model work?
Train a GAM predicting contact rate on POLYMOD data
Simplified:
How does the model work (really)?
(simplified)
mgcv::bam(
formula = contacts ~
# Prem method did a post-hoc smoothing
# abs(age_from - age_to)
s(gam_age_offdiag) +
# abs(age_from - age_to)^2
s(gam_age_offdiag_2) +
# abs(age_from * age_to)
s(gam_age_diag_prod) +
# abs(age_from + age_to)
s(gam_age_diag_sum) +
# pmax(age_from, age_to)
s(gam_age_pmax) +
# pmin(age_from, age_to)
s(gam_age_pmin) +
school_probability +
work_probability,
family = stats::poisson,
# NOTE: the offset of participants allows us to get the rate per person
offset = log(participants),
data = .
)How does the model work (really really)?
- Here’s the link to the code
Some Prem et al Limitations
Although they provided the code:
It was code not written for reuse (code vs software)
- No interface on getting matrix for a specified region.
- Unclear which code matched which methods
Australia was in original 155 countries, not in the new updated 177!
This is the last slide.

njtierney.github.io/talk-spark-spectrum-2023/